Carla A. Rudder, PMP

Data-driven professional with a strong background in mathematics, statistics, data analysis, and education. I use data to find patterns and insights, drive innovation, and create meaningful change.
Technical Skills: Excel, SQL, Python, R, Power BI, Tableau, and Certified Scrum Master
Applied Data Science & ML - MIT
BSc Applied Mathematics
MSc Mathematics Ed (Financial Math)
Ph.D. Mathematics Ed (Problem-Solving)
View My LinkedIn Profile
Loan Default Prediction
Problem Definition
The Context:
-
A major proportion of retail bank profit comes from interests in the form of home loans. These loans are borrowed by regular income/high-earning customers. Banks are most fearful of defaulters, as bad loans (NPA) usually eat up a major chunk of their profits. Therefore, it is important for banks to be judicious while approving loans for their customer base.
-
The approval process for the loans is multifaceted. Through this process, the bank tries to check the creditworthiness of the applicant on the basis of a manual study of various aspects of the application. The entire process is not only effort-intensive but also prone to wrong judgment/approval owing to human error and biases.
-
There have been attempts by many banks to automate this process by using heuristics. But with the advent of data science and machine learning, the focus has shifted to building machines that can learn this approval process and make it free of biases and more efficient. At the same time, one important thing to keep in mind is to make sure that the machine does not learn the biases that previously crept in because of the human approval process.
-
A bank’s consumer credit department aims to simplify the decision-making process for home equity lines of credit to be accepted. To do this, they will adopt the Equal Credit Opportunity Act’s guidelines to establish an empirically derived and statistically sound model for credit scoring. The model will be based on the data obtained via the existing loan underwriting process from recent applicants who have been given credit. The model will be built from predictive modeling techniques, but the model created must be interpretable enough to provide a justification for any adverse behavior (rejections).
The objective:
- Analyze and build a classification model to predict clients who are likely to default on their loans and
- Identify important features to consider while approving a loan.
- Give recommendations to the bank on the profile of the persons who are likely to default on their loan
The key questions:
- What are the key questions that need to be answered?
- What was the average number of years at the job for the delinquent clients
- What were the top 3 Jobs of the delinquent clients
- What are the lengths of the top 5 delinquent lines of credit
- Which reason for the loan request, Home Improvement or Debt consolidation has the most delinquencies
The problem formulation:
- What is it that we are trying to solve using data science?
- Using a classification model the bank should be able to predict which features or combination of features of a customer would likely default on their loan.
Data Description:
The Home Equity dataset (HMEQ) contains baseline and loan performance information for 5,960 recent home equity loans. The target (BAD) is a binary variable that indicates whether an applicant has ultimately defaulted or has been severely delinquent. This adverse outcome occurred in 1,189 cases (20 percent). 12 input variables were registered for each applicant.
- BAD: 1 = Client defaulted on loan, 0 = loan repaid
- LOAN: Amount of loan approved.
- MORTDUE: Amount due on the existing mortgage.
- VALUE: Current value of the property.
- REASON: Reason for the loan request. (HomeImp = home improvement, DebtCon= debt consolidation which means taking out a new loan to pay off other liabilities and consumer debts)
- JOB: The type of job that the loan applicant has such as manager, self, etc.
- YOJ: Years at present job.
- DEROG: Number of major derogatory reports (which indicate serious delinquency or late payments).
- DELINQ: Number of delinquent credit lines (a line of credit becomes delinquent when a borrower does not make the minimum required payments 30 to 60 days past the day on which the payments were due).
- CLAGE: Age of the oldest credit line in months.
- NINQ: Number of recent credit inquiries.
- CLNO: Number of existing credit lines.
- DEBTINC: Debt-to-income ratio (all your monthly debt payments divided by your gross monthly income. This number is one way lenders measure your ability to manage the monthly payments to repay the money you plan to borrow.
Import the necessary libraries and Data
import warnings
warnings.filterwarnings("ignore")
import sys
!{sys.executable} -m pip install missingno
!{sys.executable} -m pip install plotnine
!{sys.executable} -m pip install --upgrade matplotlib
import warnings
warnings.filterwarnings("ignore")
# Libraries for data manipulation and visualization
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import missingno as msno
import seaborn as sns
sns.set_theme()
from plotnine import ggplot, aes, geom_point, geom_bar, labs, theme, theme_bw, element_text, element_blank, scale_fill_manual
# For training and testing the data
from sklearn.model_selection import train_test_split
# Algorithms to use
from sklearn.tree import DecisionTreeClassifier
from sklearn import tree
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import BaggingClassifier
# Metrics to evaluate the model
from sklearn.metrics import confusion_matrix, classification_report, f1_score, recall_score, precision_score
from sklearn import metrics
import scipy.stats as stats
from sklearn.linear_model import LogisticRegression
# For hyperparameter tuning
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import accuracy_score
Data Overview
- Reading the dataset
- Understanding the shape of the dataset
- Checking the data types
- Checking for missing values
- Checking for duplicated values
Load the data
- Create a copy to use for data manipulation
# Load the data - original data set
client_data = pd.read_csv('hmeq.csv')
# create a copy of the data set to work with
df = client_data.copy()
Data Overview
- Observations
- Sanity checks
- View the first and last five rows of the data
# look at the first rows of the data
df.head()
| BAD | LOAN | MORTDUE | VALUE | REASON | JOB | YOJ | DEROG | DELINQ | CLAGE | NINQ | CLNO | DEBTINC | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 1100 | 25860.0 | 39025.0 | HomeImp | Other | 10.5 | 0.0 | 0.0 | 94.366667 | 1.0 | 9.0 | NaN |
| 1 | 1 | 1300 | 70053.0 | 68400.0 | HomeImp | Other | 7.0 | 0.0 | 2.0 | 121.833333 | 0.0 | 14.0 | NaN |
| 2 | 1 | 1500 | 13500.0 | 16700.0 | HomeImp | Other | 4.0 | 0.0 | 0.0 | 149.466667 | 1.0 | 10.0 | NaN |
| 3 | 1 | 1500 | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN |
| 4 | 0 | 1700 | 97800.0 | 112000.0 | HomeImp | Office | 3.0 | 0.0 | 0.0 | 93.333333 | 0.0 | 14.0 | NaN |
# check the data last five rows
df.tail()
| BAD | LOAN | MORTDUE | VALUE | REASON | JOB | YOJ | DEROG | DELINQ | CLAGE | NINQ | CLNO | DEBTINC | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 5955 | 0 | 88900 | 57264.0 | 90185.0 | DebtCon | Other | 16.0 | 0.0 | 0.0 | 221.808718 | 0.0 | 16.0 | 36.112347 |
| 5956 | 0 | 89000 | 54576.0 | 92937.0 | DebtCon | Other | 16.0 | 0.0 | 0.0 | 208.692070 | 0.0 | 15.0 | 35.859971 |
| 5957 | 0 | 89200 | 54045.0 | 92924.0 | DebtCon | Other | 15.0 | 0.0 | 0.0 | 212.279697 | 0.0 | 15.0 | 35.556590 |
| 5958 | 0 | 89800 | 50370.0 | 91861.0 | DebtCon | Other | 14.0 | 0.0 | 0.0 | 213.892709 | 0.0 | 16.0 | 34.340882 |
| 5959 | 0 | 89900 | 48811.0 | 88934.0 | DebtCon | Other | 15.0 | 0.0 | 0.0 | 219.601002 | 0.0 | 16.0 | 34.571519 |
df.shape
Observations (5960, 13)
df.info()
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 5960 entries, 0 to 5959
Data columns (total 13 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 BAD 5960 non-null int64
1 LOAN 5960 non-null int64
2 MORTDUE 5442 non-null float64
3 VALUE 5848 non-null float64
4 REASON 5708 non-null object
5 JOB 5681 non-null object
6 YOJ 5445 non-null float64
7 DEROG 5252 non-null float64
8 DELINQ 5380 non-null float64
9 CLAGE 5652 non-null float64
10 NINQ 5450 non-null float64
11 CLNO 5738 non-null float64
12 DEBTINC 4693 non-null float64
dtypes: float64(9), int64(2), object(2)
memory usage: 605.4+ KB
Observations
- There are 5960 rows and 13 columns
- The data types for the features are:
- 2 objects
REASONandJOB- Will create dummy variables for these two - 2 integers
BADandLOAN- These have all 5960 values. No NULL values - 9 floats for the remaining features
Missing Values
- Determine the number of missing values for each feature
# Count the number of missing values in each column
missing_values_count = df.isnull().sum()
# Print the number of missing values in each column
print("Number of Missing Values in Each Column:")
print(missing_values_count)
Number of Missing Values in Each Column:
BAD 0
LOAN 0
MORTDUE 518
VALUE 112
REASON 252
JOB 279
YOJ 515
DEROG 708
DELINQ 580
CLAGE 308
NINQ 510
CLNO 222
DEBTINC 1267
dtype: int64
- Percentage of missing values
# Find the percentage of missing values in each column
percent_missing = (missing_values_count/5960)*100
#print the percentage of missing values in each column
print("Percent of Missing Values in Each Column:")
print(percent_missing)
Percent of Missing Values in Each Column:
BAD 0.000000
LOAN 0.000000
MORTDUE 8.691275
VALUE 1.879195
REASON 4.228188
JOB 4.681208
YOJ 8.640940
DEROG 11.879195
DELINQ 9.731544
CLAGE 5.167785
NINQ 8.557047
CLNO 3.724832
DEBTINC 21.258389
dtype: float64
Observations
DEBTINChas the highest percent of missing values with 21%- All other features have <= 11.8% missing values
#Visulaize the number of missing values in each row
# Visualize missing values using missingno with custom colors
# Visualize missing values using missingno with custom colors
msno.matrix(df, figsize=(10, 5), color=(0.2, 0.7, 0.2), sparkline=False)
# Add a legend with custom color patches
present_patch = mpatches.Patch(color=(0.2, 0.7, 0.2), label='Present')
missing_patch = mpatches.Patch(color='white', label='Missing')
plt.legend(handles=[present_patch, missing_patch], title='Legend', bbox_to_anchor=(1.05, 1), loc='upper left')
# Show the plot
plt.show()
Visualizations of Missing Values
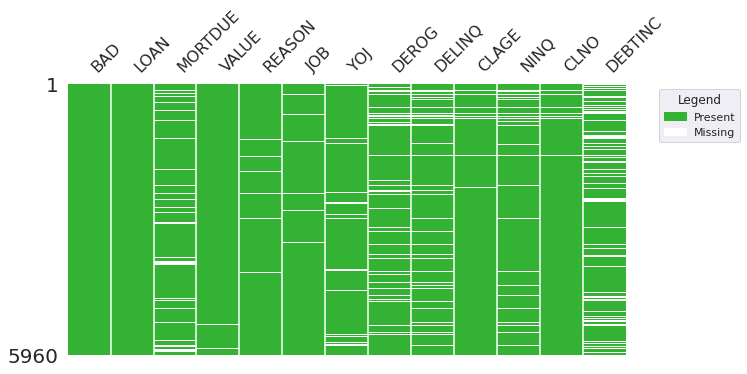
# Convert object columns to a categorical (factor)
df['BAD'] = df['BAD'].astype('category')
df['REASON'] = df['REASON'].astype('category')
df['JOB'] = df['JOB'].astype('category')
# see the data types again
df.info()
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 5960 entries, 0 to 5959
Data columns (total 13 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 BAD 5960 non-null category
1 LOAN 5960 non-null int64
2 MORTDUE 5442 non-null float64
3 VALUE 5848 non-null float64
4 REASON 5708 non-null category
5 JOB 5681 non-null category
6 YOJ 5445 non-null float64
7 DEROG 5252 non-null float64
8 DELINQ 5380 non-null float64
9 CLAGE 5652 non-null float64
10 NINQ 5450 non-null float64
11 CLNO 5738 non-null float64
12 DEBTINC 4693 non-null float64
dtypes: category(3), float64(9), int64(1)
memory usage: 483.7 KB
Summary Statistics
df.describe()
| LOAN | MORTDUE | VALUE | YOJ | DEROG | DELINQ | CLAGE | NINQ | CLNO | DEBTINC | |
|---|---|---|---|---|---|---|---|---|---|---|
| count | 5960.000000 | 5442.000000 | 5848.000000 | 5445.000000 | 5252.000000 | 5380.000000 | 5652.000000 | 5450.000000 | 5738.000000 | 4693.000000 |
| mean | 18607.969799 | 73760.817200 | 101776.048741 | 8.922268 | 0.254570 | 0.449442 | 179.766275 | 1.186055 | 21.296096 | 33.779915 |
| std | 11207.480417 | 44457.609458 | 57385.775334 | 7.573982 | 0.846047 | 1.127266 | 85.810092 | 1.728675 | 10.138933 | 8.601746 |
| min | 1100.000000 | 2063.000000 | 8000.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.524499 |
| 25% | 11100.000000 | 46276.000000 | 66075.500000 | 3.000000 | 0.000000 | 0.000000 | 115.116702 | 0.000000 | 15.000000 | 29.140031 |
| 50% | 16300.000000 | 65019.000000 | 89235.500000 | 7.000000 | 0.000000 | 0.000000 | 173.466667 | 1.000000 | 20.000000 | 34.818262 |
| 75% | 23300.000000 | 91488.000000 | 119824.250000 | 13.000000 | 0.000000 | 0.000000 | 231.562278 | 2.000000 | 26.000000 | 39.003141 |
| max | 89900.000000 | 399550.000000 | 855909.000000 | 41.000000 | 10.000000 | 15.000000 | 1168.233561 | 17.000000 | 71.000000 | 203.312149 |
Observations from Summary Statistics
- The average
LOANamount is 18607.97 .The minimum loan amount is 1100 and maximum amount is 89900. -
There is a large gap between the 75th percentile and maximum, perhaps indicating outliers.
- The average
MORTDUEis 73760.81. The minimum is 2063 and maximum amount is 3995500 -
There is a large gap between the 75th percentile and maximum, perhaps indicating outliers
-
The average
VALUEis 101776.05. The minimum is 8000 and maximum amount is 855909 - The average
YOJis 8.9. The minimum is 0 and maximum time is 41 -
There is a large gap between the 75th percentile and maximum, perhaps indicating outliers
-
The average
DEROGis 0.25. The minimum is 0 and maximum number is 10 -
The average
DELINQis 0.45. The minimum is 0 and maximum number is 15 - The average
CLAGEis 179.8. The minimum is 0 and maximum time is 1168 -
There is a large gap between the 75th percentile and maximum, perhaps indicating outliers (having a line of credit for 1168 months seems unlikely)
- The average
NINQis 1.19. The minimum is 0 and maximum number is 17 -
There is a large gap between the 75th percentile and maximum, perhaps indicating outliers
- The average
CLNOis 21.3. The minimum is 0 and maximum number is 71 -
There is a large gap between the 75th percentile and maximum, perhaps indicating outliers
- The average
DEBTINCis 33.78. The minimum is 0.52 and maximum number is 203 - There is a large gap between the 75th percentile and maximum, perhaps indicating outliers
# Check summary for categorical data
df.describe(include=['category'])
| BAD | REASON | JOB | |
|---|---|---|---|
| count | 5960 | 5708 | 5681 |
| unique | 2 | 2 | 6 |
| top | 0 | DebtCon | Other |
| freq | 4771 | 3928 | 2388 |
# Checking the count of unique values in each categorical column
cols_cat= df.select_dtypes(['category'])
for i in cols_cat.columns:
print('Unique values in',i, 'are :')
print(df[i].value_counts(normalize = True))
print('*'*50)
Unique values in BAD are :
BAD
0 0.800503
1 0.199497
Name: proportion, dtype: float64
**************************************************
Unique values in REASON are :
REASON
DebtCon 0.688157
HomeImp 0.311843
Name: proportion, dtype: float64
**************************************************
Unique values in JOB are :
JOB
Other 0.420349
ProfExe 0.224608
Office 0.166872
Mgr 0.135011
Self 0.033973
Sales 0.019187
Name: proportion, dtype: float64
**************************************************
#create a bar chart to determine the number of customers which have defaulted on their loan (1)
plt.figure(figsize = (10, 6))
ax = sns.countplot(x = 'BAD', data = df)
# Place the exact count on the top of the bar for each category using annotate
for p in ax.patches:
ax.annotate('{:.1f}'.format(p.get_height()), (p.get_x(), p.get_height()))
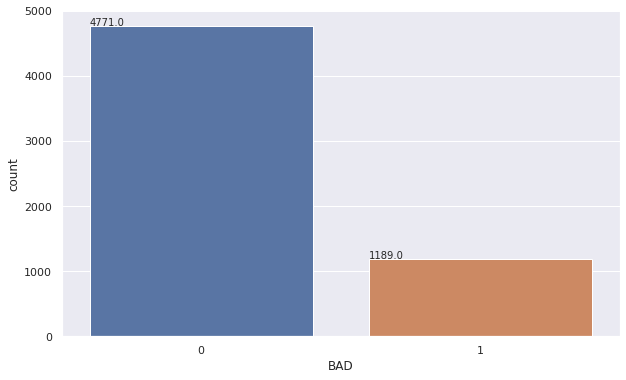
Observations for Categorical Summary
BADhas 2 options 0 (repaid) or 1(default).-
0 has the highest frequency with 4771 (80%) customers having repaid their loans and are not in default.
REASONhas two options DebtCon (Debt consolidation) or HomeImp (Home Improvement).- DebtCon has the highest frequency with 3928 (69%).
-
However there are only 5708 responses therefore there are some missing values.
JOBhas 6 options with Other having the highest frequency with 2388 (42%).- However there are only 5681 responses therefore there are some missing values.
Exploratory Data Analysis (EDA) and Visualization
- EDA is an important part of any project involving data.
- It is important to investigate and understand the data better before building a model with it.
- A few questions have been mentioned below which will help you approach the analysis in the right manner and generate insights from the data.
- A thorough analysis of the data, in addition to the questions mentioned below, should be done.
Leading Questions:
- What is the range of values for the loan amount variable “LOAN”?
- How does the distribution of years at present job “YOJ” vary across the dataset?
- How many unique categories are there in the REASON variable?
- What is the most common category in the JOB variable?
- Is there a relationship between the REASON variable and the proportion of applicants who defaulted on their loan?
- Do applicants who default have a significantly different loan amount compared to those who repay their loan?
- Is there a correlation between the value of the property and the loan default rate?
- Do applicants who default have a significantly different mortgage amount compared to those who repay their loan?
Univariate Analysis
Distributions and Outliers
Create count plots to identify the distribution of the data
Create box plots to determine if there are any outliers in the data.
#create histograms and box plots to visualize data to identify the distribution and outliers
for col in ['LOAN', 'MORTDUE', 'VALUE', 'YOJ', 'DEROG', 'DELINQ', 'CLAGE', 'NINQ', 'CLNO', 'DEBTINC']:
print(col)
print('The skew is:', round(df[col].skew(), 2))
plt.figure(figsize=(20, 4))
# histogram
plt.subplot(1, 2, 1)
plt.hist(df[col], bins=10)
plt.ylabel('count')
# Add mean and median to the histogram
mean_val = df[col].mean()
median_val = df[col].median()
plt.axvline(mean_val, color='r', linestyle='dashed', linewidth=2, label='Mean')
plt.axvline(median_val, color='g', linestyle='dashed', linewidth=2, label='Median')
plt.legend()
# box plot
plt.subplot(1, 2, 2)
sns.boxplot(df[col])
plt.show()
LOAN
The skew is: 2.02 <img src="cust_loan_pred/output_36_0.png?raw=true"/>
MORTDUE
The skew is: 1.81 <img src="cust_loan_pred/output_36_3.png?raw=true"/>
VALUE
The skew is: 3.05 <img src="cust_loan_pred/output_36_5.png?raw=true"/>
YOJ
The skew is: 0.99 <img src="cust_loan_pred/output_36_7.png?raw=true"/>
DEROG
The skew is: 5.32 <img src="cust_loan_pred/output_36_9.png?raw=true"/>
DELINQ
The skew is: 4.02 <img src="cust_loan_pred/output_36_11.png?raw=true"/>
CLAGE
The skew is: 1.34 <img src="cust_loan_pred/output_36_13.png?raw=true"/>
NINQ
The skew is: 2.62 <img src="cust_loan_pred/output_36_15.png?raw=true"/>
CLNO
The skew is: 0.78 <img src="cust_loan_pred/output_36_17.png?raw=true"/>
DEBTINC
The skew is: 2.85 <img src="cust_loan_pred/output_36_19.png?raw=true"/>
Observations
LOAN- Positive Skew (2.02) with a range from 0 to 90000 and there are outliers present on the box plot.-
Due to the low number of loans above the 75th% these should remain. The bank is only going above that in a few cases.
MORTDUE- Positive Skew (1.9) with a range from 0 to 275000 and there are outliers present on the box plot-
Considering the number of outliers outside of the 75th percentile, it is likely this is not an error.
-
VALUE- Positive Skew (3.08) with a range from 0 to 300000 and there are outliers present on the box plot YOJ- Positive Skew (1.03) with a range from 0 to 40 and there are outliers present on the box plot-
Even though there are only a few outliers it follows the trend of the data and is likely not an outlier.
DEROG- Positive Skew (5.67) with a range from 0 to 8 and there are outliers present on the box plot-
hard to say if these are outliers or just special cases.
DELINQ- Positive Skew (4.23) with a range from 0 to 7.5 and there are outliers present on the box plot-
hard to say if these are outliers or just special cases.
CLAGE- Positive Skew (1.38) with a range from 0 to 700 and there are outliers present on the box plot-
Given the spread of data points it is highly likely that the 2 max outliers are errors. Should remove them as to not contaminate the data.
NINQ- Positive Skew (2.74) with a range from 0 to 12 and there are outliers present on the box plot.-
Looks more like special cases, rather than outliers, as it follows the curve of the data.
-
CLNO- Positive Skew (2.02) with a range from 0 to 70 and there are outliers present on the box plot DEBTINC- Positive Skew (3.21) with a range from 0 to 75 and there are outliers present on the box plot. IT is not likely that a person has a Debt to income ratio greater than 200. The last outlier is likely an error, should remove it.
Univariate Analysis - Categorical Data
# Build barplot for REASON
plt.figure(figsize=(15, 5))
ax = sns.countplot(df["REASON"], palette='winter')
# Function to add percentage on top of each bar
def perc_on_bar(ax, data):
total = len(data)
for p in ax.patches:
percentage = '{:.1f}%'.format(100 * p.get_height() / total)
x = p.get_x() + p.get_width() / 2
y = p.get_y() + p.get_height()
ax.annotate(percentage, (x, y), ha='center')
perc_on_bar(ax, df["REASON"])
plt.show()
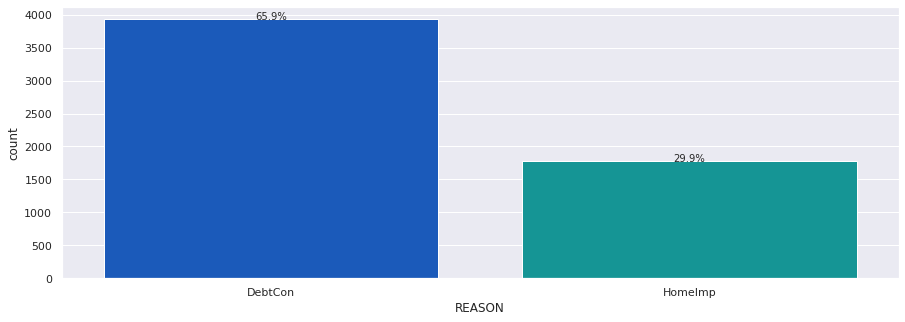
# Build barplot for JOB
plt.figure(figsize=(15, 5))
ax = sns.countplot(df["JOB"], palette='winter')
# Function to add percentage on top of each bar
def perc_on_bar(ax, data):
total = len(data)
for p in ax.patches:
percentage = '{:.1f}%'.format(100 * p.get_height() / total)
x = p.get_x() + p.get_width() / 2
y = p.get_y() + p.get_height()
ax.annotate(percentage, (x, y), ha='center')
perc_on_bar(ax, df["JOB"])
plt.show()
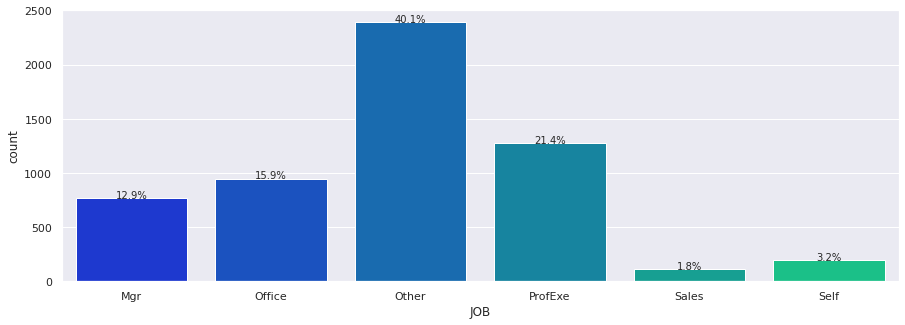
Major Observations
- There are two features that may have errors based on the outliers
CLAGEandDEBTINC
Bivariate Analysis
BAD vs LOAN
sns.boxplot(df["BAD"],df['LOAN'],palette="PuBu")
<Axes: xlabel='BAD', ylabel='LOAN'> <img src="cust_loan_pred/output_44_1.png?raw=true"/>
Observations
- Loan amount does not seem to have a great effect on defaults.
BAD vs. MORTDUE
sns.boxplot(df["BAD"],df['MORTDUE'],palette="PuBu")
<Axes: xlabel='BAD', ylabel='MORTDUE'> <img src="cust_loan_pred/output_47_1.png?raw=true"/>
Observations
- Amount due on existing mortgage does not seem to have a great impact on defaults.
BAD vs. VALUE
sns.boxplot(df["BAD"],df['VALUE'],palette="PuBu")
<Axes: xlabel='BAD', ylabel='VALUE'> <img src="cust_loan_pred/output_50_1.png?raw=true"/>
Observations
- Value of existing mortgage does not seem to have a great impact on defaults.
BAD vs. DEBTINC
sns.boxplot(df["BAD"],df['DEBTINC'],palette="PuBu")
<Axes: xlabel='BAD', ylabel='DEBTINC'> <img src="cust_loan_pred/output_53_1.png?raw=true"/>
Observations
- Greater debt to income ratio does seem to result in more defaults
Continuous Variables
VALUE and DEROG
sns.scatterplot(df["VALUE"],df['DEROG'],palette="PuBu")
<Axes: xlabel='VALUE', ylabel='DEROG'> <img src="cust_loan_pred/output_57_1.png?raw=true"/>
- The larger the loan the lower number of derogatory records.
VALUE and DELINQ
sns.scatterplot(df["VALUE"],df['DELINQ'],palette="PuBu")
<Axes: xlabel='VALUE', ylabel='DELINQ'> <img src="cust_loan_pred/output_60_1.png?raw=true"/>
- The larger the value of the loan the lower number of delinquent records
Bivariate Analysis: BAD vs Categorical Variables
The stacked bar graph allows you to look at numerical values across two categorical variables.
# Function to plot stacked bar charts for categorical columns
def stacked_plot(x):
sns.set(palette='nipy_spectral')
tab1 = pd.crosstab(x,df['BAD'],margins=True)
print(tab1)
print('-'*120)
tab = pd.crosstab(x,df['BAD'],normalize='index')
tab.plot(kind='bar',stacked=True,figsize=(10,5))
plt.legend(loc='lower left', frameon=False)
plt.legend(loc="upper left", bbox_to_anchor=(1,1))
plt.show()
Plot stacked bar plot for LOAN and JOBS
# Plot stacked bar plot for LOAN and JOB
stacked_plot(df['JOB'])
BAD 0 1 All
JOB
Mgr 588 179 767
Office 823 125 948
Other 1834 554 2388
ProfExe 1064 212 1276
Sales 71 38 109
Self 135 58 193
All 4515 1166 5681
------------------------------------------------------------------------------------------------------------------------
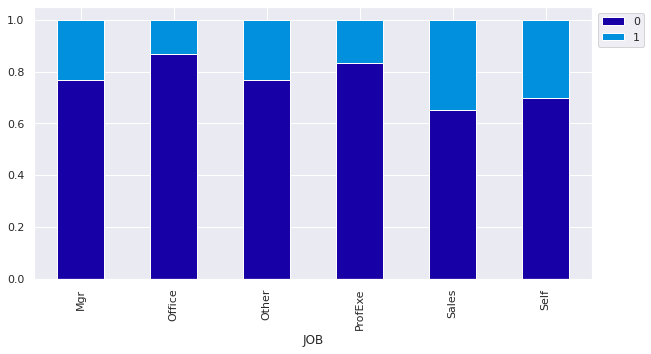
Observations
- Sales employees and Self-employed have the highest instances of defaulting on their loan.
Plot stacked bar plot for LOAN and DEROG
# Plot stacked bar plot for LOAN and DEROG
stacked_plot(df['DEROG'])
BAD 0 1 All
DEROG
0.0 3773 754 4527
1.0 266 169 435
2.0 78 82 160
3.0 15 43 58
4.0 5 18 23
5.0 8 7 15
6.0 5 10 15
7.0 0 8 8
8.0 0 6 6
9.0 0 3 3
10.0 0 2 2
All 4150 1102 5252
------------------------------------------------------------------------------------------------------------------------
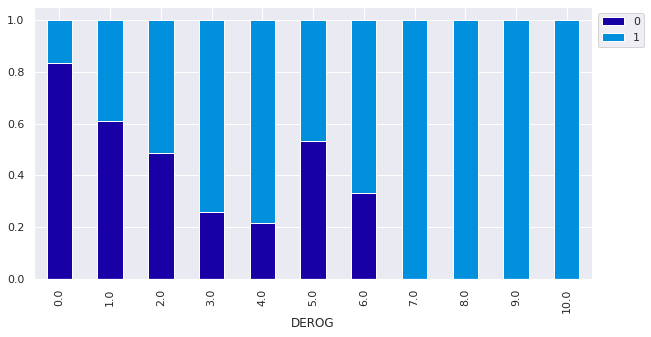
Observations
- Customers with >=7 derogatory reports all defaulted on their loans
- The number of customers who defaulted on their loans increases with the number of derogatory reports
Plot stacked bar plot for for LOAN and DELINQ
stacked_plot(df['DELINQ'])
BAD 0 1 All
DELINQ
0.0 3596 583 4179
1.0 432 222 654
2.0 138 112 250
3.0 58 71 129
4.0 32 46 78
5.0 7 31 38
6.0 0 27 27
7.0 0 13 13
8.0 0 5 5
10.0 0 2 2
11.0 0 2 2
12.0 0 1 1
13.0 0 1 1
15.0 0 1 1
All 4263 1117 5380
------------------------------------------------------------------------------------------------------------------------
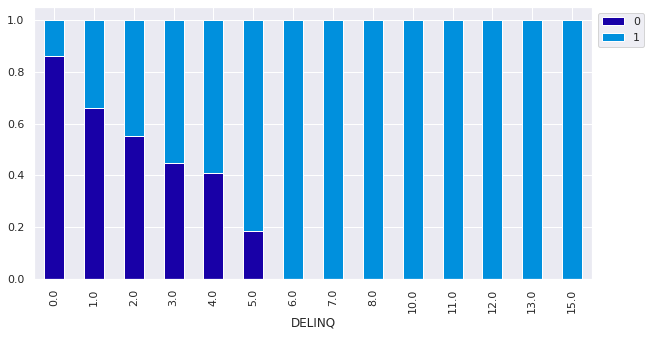
Observations
- The customers with >= 6 Delinquencies all defaulted on their loan
Multivariate Analysis
Correlation heat map
# Separating numerical variables
numerical_col = df.select_dtypes(include=np.number).columns.tolist()
# Build correlation matrix for numerical columns
corr = df[numerical_col].corr()
# plot the heatmap
plt.figure(figsize=(15,10))
sns.heatmap(corr,cmap='coolwarm',vmax=1,vmin=-1,
fmt=".2f",
annot=True,
xticklabels=corr.columns,
yticklabels=corr.columns);
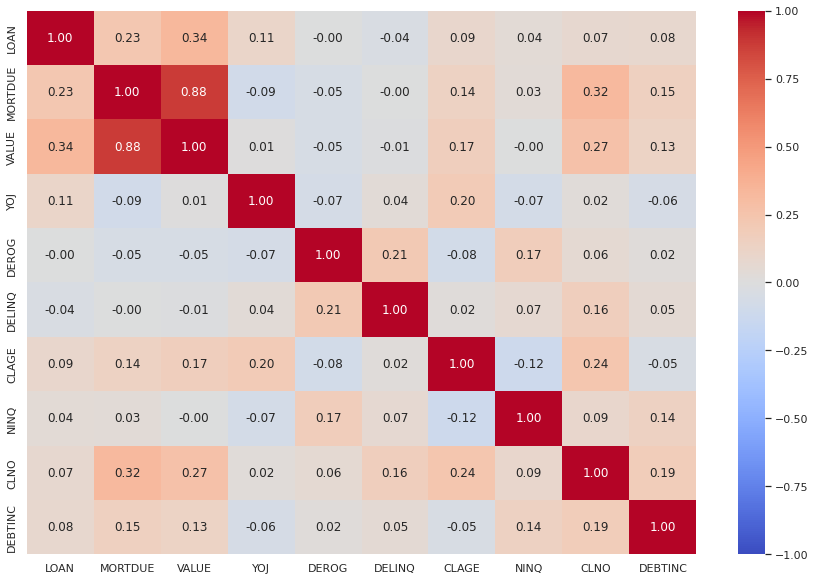
Observations
- The mortgage due and the value of the house are very closely related with 0.88
Treating Outliers
Identifying outliers
- defined a function
find_outliers_IQRwhich takes in a dataframe as an input and returns a dataframe as an output. The returned data frame contains the outliers as numerical values and others as NaN - identified the outliers: lower limit < q1 - 1.5 * IQR and higher limit > q3 + 1.5 * IQR
Replace the outliers
- if outlier is < Lower Whisker then replace outlier with Lower Whisker
- if outlier is > Upper Whisker then replace outlier with Upper Whisker
# make a copy of the original data
df_raw = df.copy()
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the LOAN
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['LOAN'])
# Replace the outliers with the whisker values
df_raw['LOAN'] = df_raw['LOAN'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'LOAN' column
num_outliers = ((df_raw['LOAN'] == Lower_Whisker) | (df_raw['LOAN'] == Upper_Whisker)).sum()
print('Number of outliers in LOAN column:', num_outliers)
print('LOAN min outlier value: ' + str(outliers.min()))
print('LOAN max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in LOAN column: 260
LOAN min outlier value: 41700
LOAN max outlier value: 89900
5704 41700
5705 41700
5706 41700
5707 41700
5708 41800
...
5955 88900
5956 89000
5957 89200
5958 89800
5959 89900
Name: LOAN, Length: 256, dtype: int64
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the LOAN
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['LOAN'])
# Replace the outliers with the whisker values
df_raw['MORTDUE'] = df_raw['MORTDUE'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'MORTDUE' column
num_outliers = ((df_raw['MORTDUE'] == Lower_Whisker) | (df_raw['MORTDUE'] == Upper_Whisker)).sum()
print('Number of outliers in MORTDUE column:', num_outliers)
print('MORTDUE min outlier value: ' + str(outliers.min()))
print('MORTDUE max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in MORTDUE column: 4355
MORTDUE min outlier value: nan
MORTDUE max outlier value: nan
Series([], Name: LOAN, dtype: int64)
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the VALUE
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['VALUE'])
# Replace the outliers with the whisker values
df_raw['VALUE'] = df_raw['VALUE'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'VALUE' column
num_outliers = ((df_raw['VALUE'] == Lower_Whisker) | (df_raw['VALUE'] == Upper_Whisker)).sum()
print('Number of outliers in VALUE column:', num_outliers)
print('VALUE min outlier value: ' + str(outliers.min()))
print('VALUE max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in VALUE column: 320
VALUE min outlier value: 200459.0
VALUE max outlier value: 855909.0
140 245300.0
194 251962.0
291 250155.0
345 245730.0
414 208910.0
...
5927 288512.0
5928 292380.0
5929 289430.0
5933 215000.0
5934 224630.0
Name: VALUE, Length: 320, dtype: float64
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the YOJ
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['YOJ'])
# Replace the outliers with the whisker values
df_raw['YOJ'] = df_raw['YOJ'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'YOJ' column
num_outliers = ((df_raw['YOJ'] == Lower_Whisker) | (df_raw['YOJ'] == Upper_Whisker)).sum()
print('Number of outliers in YOJ column:', num_outliers)
print('YOJ min outlier value: ' + str(outliers.min()))
print('YOJ max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in YOJ column: 126
YOJ min outlier value: 28.5
YOJ max outlier value: 41.0
263 31.0
407 29.0
408 29.0
412 29.0
416 31.0
...
5687 30.0
5723 30.0
5741 30.0
5747 31.0
5900 33.0
Name: YOJ, Length: 91, dtype: float64
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the DEROG
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['DEROG'])
# Replace the outliers with the whisker values
df_raw['DEROG'] = df_raw['DEROG'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'DEROG' column
num_outliers = ((df_raw['DEROG'] == Lower_Whisker) | (df_raw['DEROG'] == Upper_Whisker)).sum()
print('Number of outliers in DEROG column:', num_outliers)
print('DEROG min outlier value: ' + str(outliers.min()))
print('DEROG max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in DEROG column: 5252
DEROG min outlier value: 1.0
DEROG max outlier value: 10.0
6 3.0
16 2.0
21 1.0
28 1.0
31 2.0
...
5867 2.0
5877 1.0
5879 2.0
5881 2.0
5933 1.0
Name: DEROG, Length: 725, dtype: float64
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the DELINQ
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['DELINQ'])
# Replace the outliers with the whisker values
df_raw['DELINQ'] = df_raw['DELINQ'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'DELINQ' column
num_outliers = ((df_raw['DELINQ'] == Lower_Whisker) | (df_raw['DELINQ'] == Upper_Whisker)).sum()
print('Number of outliers in DELINQ column:', num_outliers)
print('DELINQ min outlier value: ' + str(outliers.min()))
print('DELINQ max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in DELINQ column: 5380
DELINQ min outlier value: 1.0
DELINQ max outlier value: 15.0
1 2.0
6 2.0
8 2.0
11 1.0
14 1.0
...
5939 1.0
5940 1.0
5941 1.0
5947 1.0
5954 1.0
Name: DELINQ, Length: 1201, dtype: float64
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the CLAGE
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['CLAGE'])
# Replace the outliers with the whisker values
df_raw['CLAGE'] = df_raw['CLAGE'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'CLAGE' column
num_outliers = ((df_raw['CLAGE'] == Lower_Whisker) | (df_raw['CLAGE'] == Upper_Whisker)).sum()
print('Number of outliers in CLAGE column:', num_outliers)
print('CLAGE min outlier value: ' + str(outliers.min()))
print('CLAGE max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in CLAGE column: 47
CLAGE min outlier value: 407.26116688
CLAGE max outlier value: 1168.2335609
202 417.633333
379 419.375154
411 475.800000
423 423.209624
469 421.541928
540 411.952051
712 419.272965
786 420.098164
909 411.753081
1166 419.033333
1205 421.362872
1470 440.421254
1797 427.923591
1848 436.751780
2010 407.261167
3097 1154.633333
3679 1168.233561
3787 630.033333
3856 632.103186
3868 618.735890
3928 634.461893
3976 407.585624
3980 412.014873
4040 626.297094
4059 623.456209
4098 627.702390
4143 626.771393
4153 615.133373
4238 638.275361
4262 628.158062
4264 639.058172
4357 622.355768
4390 628.981948
4402 627.662080
4417 629.095766
4431 468.866667
4470 649.747104
4535 408.187646
4536 648.328493
4538 412.020547
4656 471.887527
4670 473.813986
4682 474.027074
4931 485.945358
4990 480.356014
5044 476.728281
5060 468.178122
Name: CLAGE, dtype: float64
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the NINQ
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['NINQ'])
# Replace the outliers with the whisker values
df_raw['NINQ'] = df_raw['NINQ'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'NINQ' column
num_outliers = ((df_raw['NINQ'] == Lower_Whisker) | (df_raw['NINQ'] == Upper_Whisker)).sum()
print('Number of outliers in NINQ column:', num_outliers)
print('NINQ min outlier value: ' + str(outliers.min()))
print('NINQ max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in NINQ column: 252
NINQ min outlier value: 6.0
NINQ max outlier value: 17.0
44 14.0
175 10.0
212 10.0
273 10.0
283 9.0
...
5876 7.0
5879 7.0
5881 6.0
5930 9.0
5931 9.0
Name: NINQ, Length: 177, dtype: float64
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the CLNO
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['CLNO'])
# Replace the outliers with the whisker values
df_raw['CLNO'] = df_raw['CLNO'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'NINQ' column
num_outliers = ((df_raw['CLNO'] == Lower_Whisker) | (df_raw['CLNO'] == Upper_Whisker)).sum()
print('Number of outliers in CLNO column:', num_outliers)
print('CLNO min outlier value: ' + str(outliers.min()))
print('CLNO max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in CLNO column: 219
CLNO min outlier value: 43.0
CLNO max outlier value: 71.0
31 45.0
202 50.0
316 47.0
340 47.0
360 47.0
...
5617 64.0
5663 52.0
5773 46.0
5783 48.0
5833 47.0
Name: CLNO, Length: 219, dtype: float64
# Define the function to identify outliers
def find_outliers_IQR(df_raw):
Q1 = df_raw.quantile(0.25)
Q3 = df_raw.quantile(0.75)
IQR = Q3 - Q1
Lower_Whisker = Q1 - 1.5 * IQR
Upper_Whisker = Q3 + 1.5 * IQR
outliers = df_raw[((df_raw < Lower_Whisker) | (df_raw > Upper_Whisker))]
return outliers, Lower_Whisker, Upper_Whisker
# Identifying the outliers for the DEBTINC
outliers, Lower_Whisker, Upper_Whisker = find_outliers_IQR(df_raw['DEBTINC'])
# Replace the outliers with the whisker values
df_raw['DEBTINC'] = df_raw['DEBTINC'].clip(lower=Lower_Whisker, upper=Upper_Whisker)
# Count the number of outliers in the 'DEBTINC' column
num_outliers = ((df_raw['DEBTINC'] == Lower_Whisker) | (df_raw['DEBTINC'] == Upper_Whisker)).sum()
print('Number of outliers in DEBTINC column:', num_outliers)
print('DEBTINC min outlier value: ' + str(outliers.min()))
print('DEBTINC max outlier value: ' + str(outliers.max()))
outliers
Number of outliers in DEBTINC column: 94
DEBTINC min outlier value: 0.5244992154
DEBTINC max outlier value: 203.31214869
17 3.711312
57 0.524499
118 11.652739
216 72.670433
220 12.043671
...
5731 13.417822
5732 14.042012
5734 13.312356
5826 3.342570
5912 144.189001
Name: DEBTINC, Length: 94, dtype: float64
Checking that outliers have been removed
# Function that will create boxplot and histogram for any input numerical variable.
# This function takes the numerical column as the input and return the boxplots and histograms for the variable.
def histogram_boxplot(feature, figsize=(15,10), bins = None):
""" Boxplot and histogram combined
feature: 1-d feature array
figsize: size of fig (default (9,8))
bins: number of bins (default None / auto)
"""
f2, (ax_box2, ax_hist2) = plt.subplots(nrows = 2, # Number of rows of the subplot grid= 2
sharex = True, # x-axis will be shared among all subplots
gridspec_kw = {"height_ratios": (.25, .75)},
figsize = figsize
) # creating the 2 subplots
sns.boxplot(feature, ax=ax_box2, showmeans=True, color='violet') # boxplot will be created and a star will indicate the mean value of the column
sns.distplot(feature, kde=F, ax=ax_hist2, bins=bins,palette="winter") if bins else sns.distplot(feature, kde=False, ax=ax_hist2) # For histogram
ax_hist2.axvline(np.mean(feature), color='green', linestyle='--') # Add mean to the histogram
ax_hist2.axvline(np.median(feature), color='black', linestyle='-') # Add median to the histogram
histogram_boxplot(df['VALUE'])
histogram_boxplot(df_raw['VALUE'])
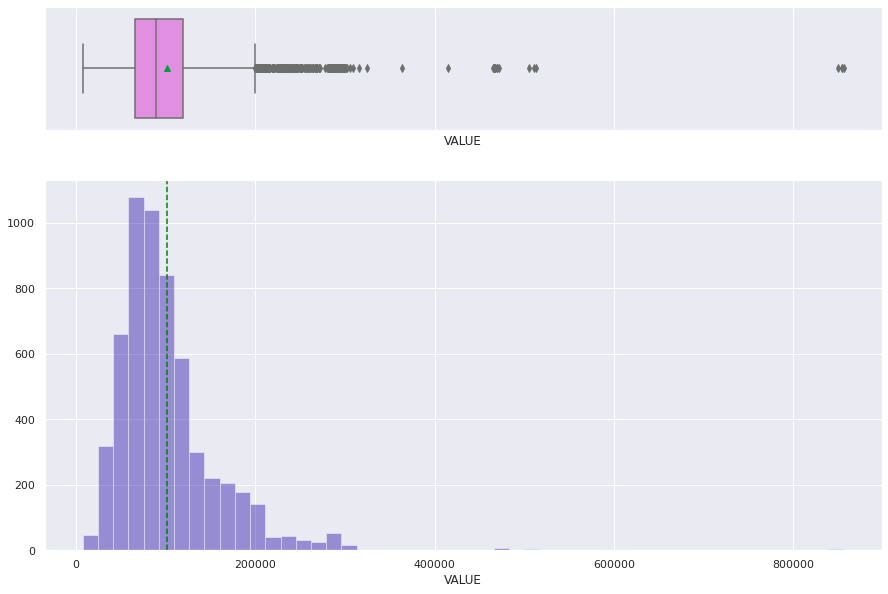
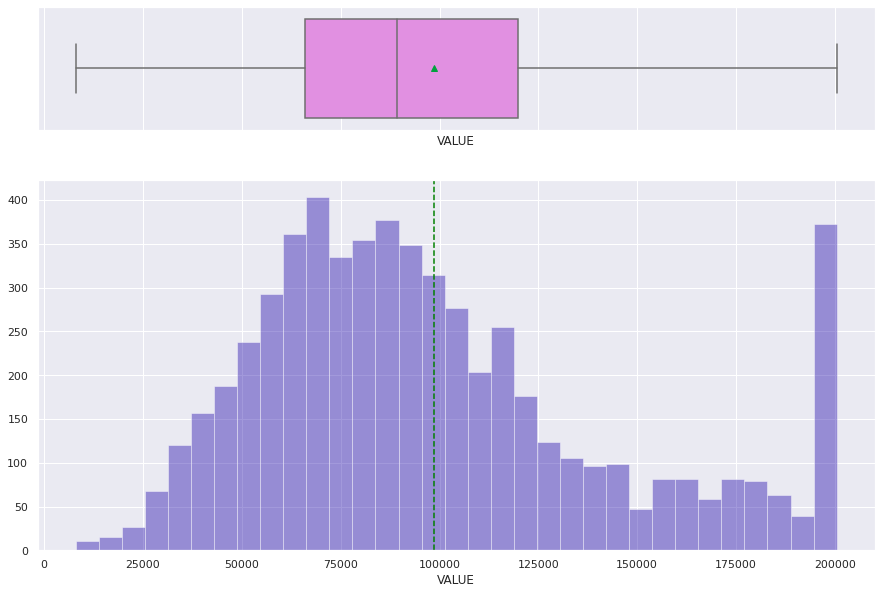
Treating Missing Values
Adding new columns in the dataset for each column which has missing values
# Will look at each row to first determine the number of missing values in each row.
# create a column to give the number of missing values in the row.
# Create a column to say 'keep' if number of missing values os <7 and 'del' if >=7 values
# Create a column to count the number of missing/null values in each row
df_raw['num_null'] = df_raw.isnull().sum(axis=1)
# Create a column to label rows as 'has_null' if num_null >= 1, otherwise 'comp' for complete data
df_raw['label'] = df_raw['num_null'].apply(lambda x: 'has_null' if x >= 1 else 'comp')
# look at top 5 rows of the data frame with the additional columns
df_raw.head()
| BAD | LOAN | MORTDUE | VALUE | REASON | JOB | YOJ | DEROG | DELINQ | CLAGE | NINQ | CLNO | DEBTINC | num_null | label | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 1100 | 25860.0 | 39025.0 | HomeImp | Other | 10.5 | 0.0 | 0.0 | 94.366667 | 1.0 | 9.0 | NaN | 1 | has_null |
| 1 | 1 | 1300 | 41600.0 | 68400.0 | HomeImp | Other | 7.0 | 0.0 | 0.0 | 121.833333 | 0.0 | 14.0 | NaN | 1 | has_null |
| 2 | 1 | 1500 | 13500.0 | 16700.0 | HomeImp | Other | 4.0 | 0.0 | 0.0 | 149.466667 | 1.0 | 10.0 | NaN | 1 | has_null |
| 3 | 1 | 1500 | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | NaN | 11 | has_null |
| 4 | 0 | 1700 | 41600.0 | 112000.0 | HomeImp | Office | 3.0 | 0.0 | 0.0 | 93.333333 | 0.0 | 14.0 | NaN | 1 | has_null |
# Count the number of 'del' in the 'label' column
num_del = (df_raw['label'] == 'has_null').sum()
# Print the number of 'del' in the 'label' column
print("Number of rows with missing values are:", num_del)
Number of rows with missing values are: 2596
Actions Taken
- Counted the number of missing/Null values in each row
- Created a column
num_nullto determine the number of missing/Null values in the row - Created a column
labelto say ‘comp’ or “has_null’. - There are 3364 rows which were complete. There are 2596 rows which contain missing data.
2596 rows which have missing values
- There are too many rows that have missing data to delete them all.
- Will replace the missing numerical data with the median of the column. and the category with mode of the column.
# Find columns of data type category
df_obj = df_raw.select_dtypes(include=['category']).columns
# Replace missing values in category columns with the mode of the column
for col in df_obj:
mode_value = df_raw[col].mode()[0] # Extract the mode value
df_raw[col] = df_raw[col].fillna(mode_value) # Fill NaN with mode value
Replace the missing values in the numerical columns with the mean of the column
# Find columns of data type int64 and float64
numeric_cols = df_raw.select_dtypes(include=['int64', 'float64']).columns
# Replace missing values in numeric columns with the average (median) of the column
for col in numeric_cols:
df_raw[col] = df_raw[col].fillna(df_raw[col].median())
# recehck the number of missing values
df_raw.info()
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 5960 entries, 0 to 5959
Data columns (total 15 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 BAD 5960 non-null category
1 LOAN 5960 non-null int64
2 MORTDUE 5960 non-null float64
3 VALUE 5960 non-null float64
4 REASON 5960 non-null category
5 JOB 5960 non-null category
6 YOJ 5960 non-null float64
7 DEROG 5960 non-null float64
8 DELINQ 5960 non-null float64
9 CLAGE 5960 non-null float64
10 NINQ 5960 non-null float64
11 CLNO 5960 non-null float64
12 DEBTINC 5960 non-null float64
13 num_null 5960 non-null int64
14 label 5960 non-null object
dtypes: category(3), float64(9), int64(2), object(1)
memory usage: 576.8+ KB
df_raw.head()
| BAD | LOAN | MORTDUE | VALUE | REASON | JOB | YOJ | DEROG | DELINQ | CLAGE | NINQ | CLNO | DEBTINC | num_null | label | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 1100 | 25860.0 | 39025.0 | HomeImp | Other | 10.5 | 0.0 | 0.0 | 94.366667 | 1.0 | 9.0 | 34.818262 | 1 | has_null |
| 1 | 1 | 1300 | 41600.0 | 68400.0 | HomeImp | Other | 7.0 | 0.0 | 0.0 | 121.833333 | 0.0 | 14.0 | 34.818262 | 1 | has_null |
| 2 | 1 | 1500 | 13500.0 | 16700.0 | HomeImp | Other | 4.0 | 0.0 | 0.0 | 149.466667 | 1.0 | 10.0 | 34.818262 | 1 | has_null |
| 3 | 1 | 1500 | 41600.0 | 89235.5 | DebtCon | Other | 7.0 | 0.0 | 0.0 | 173.466667 | 1.0 | 20.0 | 34.818262 | 11 | has_null |
| 4 | 0 | 1700 | 41600.0 | 112000.0 | HomeImp | Office | 3.0 | 0.0 | 0.0 | 93.333333 | 0.0 | 14.0 | 34.818262 | 1 | has_null |
# Drop the 'num_null' and 'label' columns from df_raw
df_raw = df_raw.drop(['num_null', 'label'], axis=1)
# check the columns have been dropped
df_raw.head()
| BAD | LOAN | MORTDUE | VALUE | REASON | JOB | YOJ | DEROG | DELINQ | CLAGE | NINQ | CLNO | DEBTINC | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 1100 | 25860.0 | 39025.0 | HomeImp | Other | 10.5 | 0.0 | 0.0 | 94.366667 | 1.0 | 9.0 | 34.818262 |
| 1 | 1 | 1300 | 41600.0 | 68400.0 | HomeImp | Other | 7.0 | 0.0 | 0.0 | 121.833333 | 0.0 | 14.0 | 34.818262 |
| 2 | 1 | 1500 | 13500.0 | 16700.0 | HomeImp | Other | 4.0 | 0.0 | 0.0 | 149.466667 | 1.0 | 10.0 | 34.818262 |
| 3 | 1 | 1500 | 41600.0 | 89235.5 | DebtCon | Other | 7.0 | 0.0 | 0.0 | 173.466667 | 1.0 | 20.0 | 34.818262 |
| 4 | 0 | 1700 | 41600.0 | 112000.0 | HomeImp | Office | 3.0 | 0.0 | 0.0 | 93.333333 | 0.0 | 14.0 | 34.818262 |
# check for multicollinearity between variables
# Select numeric columns
numeric_df = df_raw.select_dtypes(include=['number'])
# Correlation matrix (no grouping)
plt.figure(figsize=(15, 10))
sns.heatmap(numeric_df.corr().round(2), annot=True)
plt.title('Correlation matrix of data', fontsize=30)
plt.show()
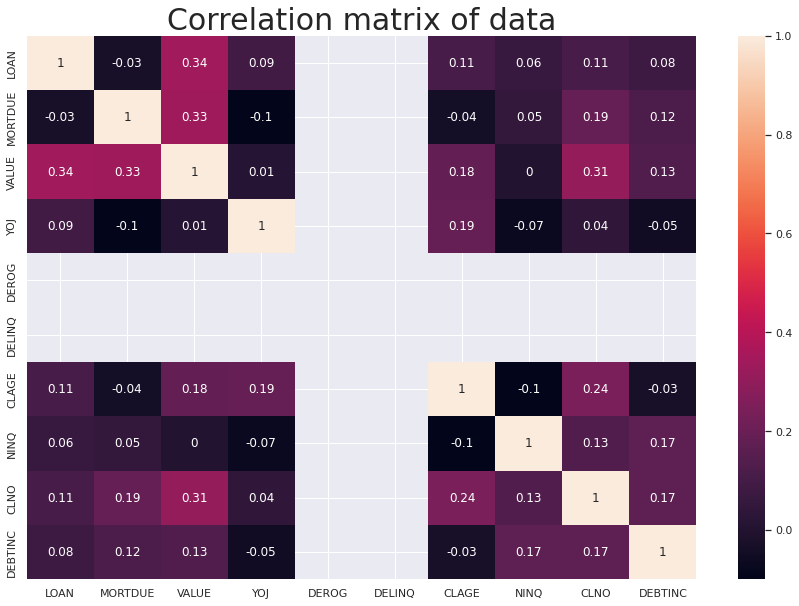
Important Insights from EDA
What are the the most important observations and insights from the data based on the EDA performed?
- Borrowers who defaulted have more delinquent credit lines and major derogatory reports than those who did not.
- Borrowers who repay their loans are credible to the bank, hence have older credit lines,
- Borrowers who repay their loans have more recent credit inquiries, and generally have more credit lines in total than default customers.
- Borrowers who repay their loans have significantly higher debt-income ratio as well as loan request amount
- Multicollinearity - or co-dependence of variables - can significantly reduce the precision of estimated beta coefficients, as a result weakening the statistical significance of regression model. This is a phenomenon we wish to avoid in our final model. However after repairing the outliers and replacing the missing / Null values this does not occur.
Model Building - Approach
- Data preparation
- Partition the data into train and test set
- Build the model
- Fit on the train data
- Tune the model
- Test the model on test set
Data Preparation
Separating the target variable from other variables
#clean data set for LG
df_lg = df_raw.copy()
# make a new copy of the clean data
df_clean = df_raw.copy()
# Drop the dependent variable from the dataframe and create the X(independent variable) matrix
X = df_clean.drop(columns = 'BAD') #make a copy called X which is a dataframe with "BAD" removed
# Create y(dependent varibale)
Y = df_clean['BAD'] #Y is a series (column) containing the "BAD" label data
# Create dummy variables for the categorical variables
X = pd.get_dummies(X, drop_first = True)
#pd.get_dummies => working on X dataframe converts all categorical variables into binary 1/0 yes/no columns.
Splitting the data into 70% train and 30% test set
# Split the data into training and test set
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = 0.30, random_state = 1)
Shape
- Check the shape of the trainng and test sets of data after manipulation
print("Shape of the training set: ", X_train.shape)
print("Shape of the test set: ", X_test.shape)
print("Percentage of classes in the training set:")
print(y_train.value_counts(normalize = True))
print("Percentage of classes in the test set:")
print(y_test.value_counts(normalize = True))
Shape of the training set: (4172, 16)
Shape of the test set: (1788, 16)
Percentage of classes in the training set:
BAD
0 0.804171
1 0.195829
Name: proportion, dtype: float64
Percentage of classes in the test set:
BAD
0 0.791946
1 0.208054
Name: proportion, dtype: float64
Logistic Regression
#Split the data into training and testing
# 70/30 split
# All rows where 'BAD' column is 1
input_ones = df_lg[df_lg['BAD'] == 1]
# All rows where 'BAD' column is 0
input_zeros = df_lg[df_lg['BAD'] == 0]
# For repeatability of sample
np.random.seed(100)
input_ones_training_rows = np.random.choice(input_ones.index, int(0.7 * len(input_ones)), replace=False)
input_zeros_training_rows = np.random.choice(input_zeros.index, int(0.7 * len(input_zeros)), replace=False)
# Pick as many 0's and 1's
training_ones = input_ones.loc[input_ones_training_rows]
training_zeros = input_zeros.loc[input_zeros_training_rows]
# Concatenate 1's and 0's
trainingData = pd.concat([training_ones, training_zeros])
# Create test data
test_ones = input_ones.drop(input_ones_training_rows)
test_zeros = input_zeros.drop(input_zeros_training_rows)
# Concatenate 1's and 0's
testData = pd.concat([test_ones, test_zeros])
#check for imbalance
bad_counts = trainingData['BAD'].value_counts()
print(bad_counts)
BAD
0 3339
1 832
Name: count, dtype: int64
# check class distrubution
class_distribution = trainingData['BAD'].value_counts(normalize=True)
print(class_distribution)
BAD
0 0.800527
1 0.199473
Name: proportion, dtype: float64
# Visualize class distribution using a bar plot
plt.figure(figsize=(8, 6))
sns.barplot(x=class_distribution.index, y=class_distribution.values)
plt.xlabel('Class (0: Good, 1: Bad)')
plt.ylabel('Proportion')
plt.title('Class Distribution')
plt.show()
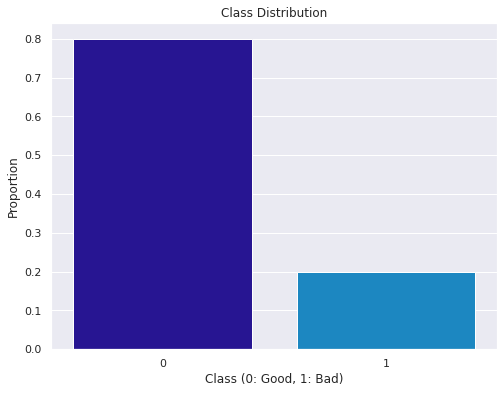
Observation
- There is a class bias
- To address class imbalance we need to use techniques like oversampling the minority class, undersampling the majority class, or using synthetic data generation techniques (e.g., SMOTE).
Undersampling
-
This method works with majority class. It reduces the number of obervations from majority class to make the data set balanced. This method is best to use when the data set is huge and reducing the number of training samples helps to improve run time an storage troubles.
-
2 types: Random and Informative
Oversampling
-
This method works with minority class. It tries to balance dataset by increasing the size of rare samples. It is used when the quantity of data is insufficient. New rare samples are generated by using e.g. repetition, bootstrapping or SMOTE (Synthetic Minority Over-Sampling Technique).
-
2 types: Random and Informative
**Resampling Training Set **
- Will use a decision tree to determine which method is best
Decision Tree often perform well on imbalanced datasets. The splitting rules that look at the class variable used in the creation of the trees, can force both classed to be addressed.
Model Evaluation Criterion
Model is attempting to find those that will default(1) on their loan, which will be our True Positive(TP), and therefore non-defaulters(0) will be our True Negative(TN)
The model can make wrong predictions as:
- Predicting an applicant will not default on a loan but, in reality, the applicant would default this is a major loss in profit for the BANK.
- Predicting an applicant will default on a loan but, in reality, the applicant would have paid it off results in the bank losing profit from the interest of that potential customer.
Which case is more important?
Predicting an applicant will not default on a loan but, in reality, the applicant would default is a major loss in profit for the BANK.
How to reduce the losses?
The bank would want recall to be maximized. The greater the recall score, the higher the chances of minimizing False Negative. In this case the false negative is predicting an applicant will not default(0), when the applicant would default(1)
That being said a high F1-Score is still preferable as that would result in more profits, as long as recall remains high.
METRIC Function
#creating metric function
def metrics_score(actual, predicted):
print(classification_report(actual, predicted))
cm = confusion_matrix(actual, predicted)
plt.figure(figsize=(8,5))
sns.heatmap(cm, annot=True, fmt='.2f', xticklabels=['Not Eligible', 'Eligible'], yticklabels=['Not Eligible', 'Eligible'])
plt.ylabel('Actual')
plt.xlabel('Predicted')
plt.show()
#Create a table to add the results of the model
results = pd.DataFrame(columns = ['Model_Name','Train_f1','Train_recall','Test_f1','Test_recall','Test_precision'])
results.head()
| Model_Name | Train_f1 | Train_recall | Test_f1 | Test_recall | Test_precision |
|---|
Decision Tree
- Encoding the data
- Creating Dummy variables
- Separating target variable from features
- Create a fit the decision tree
- Make predictions on the test model and calculate the accuracy
Build Decision tree model
#Defining Decision tree model with class weights class_weight={0: 0.2, 1: 0.8} to address data imbalance
d_tree_base = DecisionTreeClassifier(random_state = 7, class_weight = {0: 0.2, 1: 0.8})
#fitting Decision tree model
d_tree_base.fit(X_train, y_train)
DecisionTreeClassifier(class_weight={0: 0.2, 1: 0.8}, random_state=7)
Checking the performance on the train dataset
# Checking performance on the training data
y_pred_train1 = d_tree_base.predict(X_train)
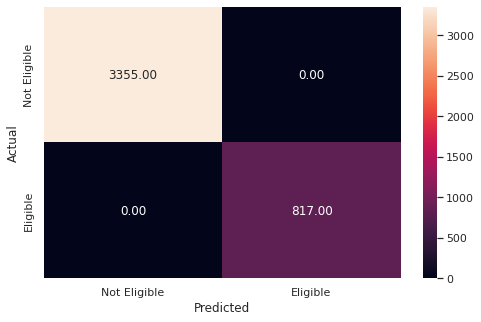
metrics_score(y_train,y_pred_train1)
precision recall f1-score support
0 1.00 1.00 1.00 3355
1 1.00 1.00 1.00 817
accuracy 1.00 4172
macro avg 1.00 1.00 1.00 4172
weighted avg 1.00 1.00 1.00 4172

Checking the performance on the test dataset
# Checking performance on the testing data
y_pred_test1 = d_tree_base.predict(X_test)
metrics_score(y_test,y_pred_test1)
precision recall f1-score support
0 0.89 0.92 0.90 1416
1 0.64 0.56 0.60 372
accuracy 0.84 1788
macro avg 0.76 0.74 0.75 1788
weighted avg 0.84 0.84 0.84 1788

Add data to the results table
# Adding the results to the table
new_row = {'Model_Name': 'd_tree_base',
'Train_f1': 100,
'Train_recall': 100,
'Test_f1': 60,
'Test_recall': 56,
'Test_precision': 64}
results = pd.concat([results, pd.DataFrame([new_row])], ignore_index=True)
# Print the updated DataFrame
print(results)
Model_Name Train_f1 Train_recall Test_f1 Test_recall Test_precision
0 d_tree_base 100 100 60 56 64
Observations
- The base model is highly overfitted with the training data out performing the testing data.
- The model has higher precision (64) to recall score (56).
- We would like the Recall Score to be higher than the Precision Score.
- Overall this is NOT a very good model.
Decision Tree - Hyperparameter Tuning
- Hyperparameter tuning is tricky in the sense that there is no direct way to calculate how a change in the hyperparameter value will reduce the loss of your model, so we usually resort to experimentation. We’ll use Grid search to perform hyperparameter tuning.
- Grid search is a tuning technique that attempts to compute the optimum values of hyperparameters.
- It is an exhaustive search that is performed on the specific parameter values of a model.
- The parameters of the estimator/model used to apply these methods are optimized by cross-validated grid-search over a parameter grid.
Criterion {“gini”, “entropy”}
The function to measure the quality of a split. Supported criteria are “gini” for the Gini impurity and “entropy” for the information gain.
max_depth
The maximum depth of the tree. If None, then nodes are expanded until all leaves are pure or until all leaves contain less than min_samples_split samples.
min_samples_leaf
The minimum number of samples is required to be at a leaf node. A split point at any depth will only be considered if it leaves at least min_samples_leaf training samples in each of the left and right branches. This may have the effect of smoothing the model, especially in regression.
You can learn about more Hyperpapameters on this link and try to tune them.
https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html
Using GridSearchCV for Hyperparameter tuning on the model
# Choose the type of classifier.
d_tree_tuned = DecisionTreeClassifier(random_state = 7, class_weight = {0: 0.2, 1: 0.8})
# Grid of parameters to choose from
parameters = {'max_depth': np.arange(2, 6), #dept [2, 3, 4, 5]
'criterion': ['gini', 'entropy'], #use both gini and entropy to measure split quality
'min_samples_leaf': [5, 10, 20, 25] #minimum number of samples to be a leaf node
}
# Type of scoring used to compare parameter combinations
scorer = metrics.make_scorer(recall_score, pos_label = 1)
# Run the grid search
grid_obj = GridSearchCV(d_tree_tuned, parameters, scoring = scorer, cv = 5) #=> chooses the best hyperparameters to use
grid_obj = grid_obj.fit(X_train, y_train)
# Set the classifier to the best combination of parameters
d_tree_tuned = grid_obj.best_estimator_
# Fit the best algorithm to the data.
d_tree_tuned.fit(X_train, y_train)
DecisionTreeClassifier(class_weight={0: 0.2, 1: 0.8}, max_depth=5,
min_samples_leaf=25, random_state=7)
Checking the performance on the training dataset
# Checking performance on the training data based on the tuned model
y_pred_train2 = d_tree_tuned.predict(X_train)
metrics_score(y_train,y_pred_train2)
precision recall f1-score support
0 0.94 0.87 0.90 3355
1 0.59 0.79 0.68 817
accuracy 0.85 4172
macro avg 0.77 0.83 0.79 4172
weighted avg 0.88 0.85 0.86 4172

Checking the performance on the test dataset
# Checking performance on the testing data based on the tuned model
y_pred_test2 = d_tree_tuned.predict(X_test)
metrics_score(y_test,y_pred_test2)
precision recall f1-score support
0 0.93 0.87 0.90 1416
1 0.60 0.75 0.67 372
accuracy 0.85 1788
macro avg 0.77 0.81 0.78 1788
weighted avg 0.86 0.85 0.85 1788

# Adding the results to the table
new_row = {'Model_Name': 'd_tree_base',
'Train_f1': 68,
'Train_recall': 79,
'Test_f1': 67,
'Test_recall': 75,
'Test_precision': 60}
results = pd.concat([results, pd.DataFrame([new_row])], ignore_index=True)
# Print the updated DataFrame
print(results)
Model_Name Train_f1 Train_recall Test_f1 Test_recall Test_precision
0 d_tree_base 100 100 60 56 64
1 d_tree_base 68 79 67 75 60
Observations
- The tuned model is slightly overfitted with a training f1-score of 68 and a testing f1-score of 67.
- The model is performing higher on recall (75) over precision (60), which is what we need to minimize false negatives.
- However, the model is overall significantly less accurate.
- Therefore we need to try and maintain these ratios while improving overall accuracy.
Plotting the Decision Tree
# Plot the decision tree and analyze it to build the decision rule
features = list(X.columns)
plt.figure(figsize = (20, 20))
tree.plot_tree(d_tree_tuned, feature_names = features, filled = True, fontsize = 12, node_ids = True, class_names = True)
plt.show()
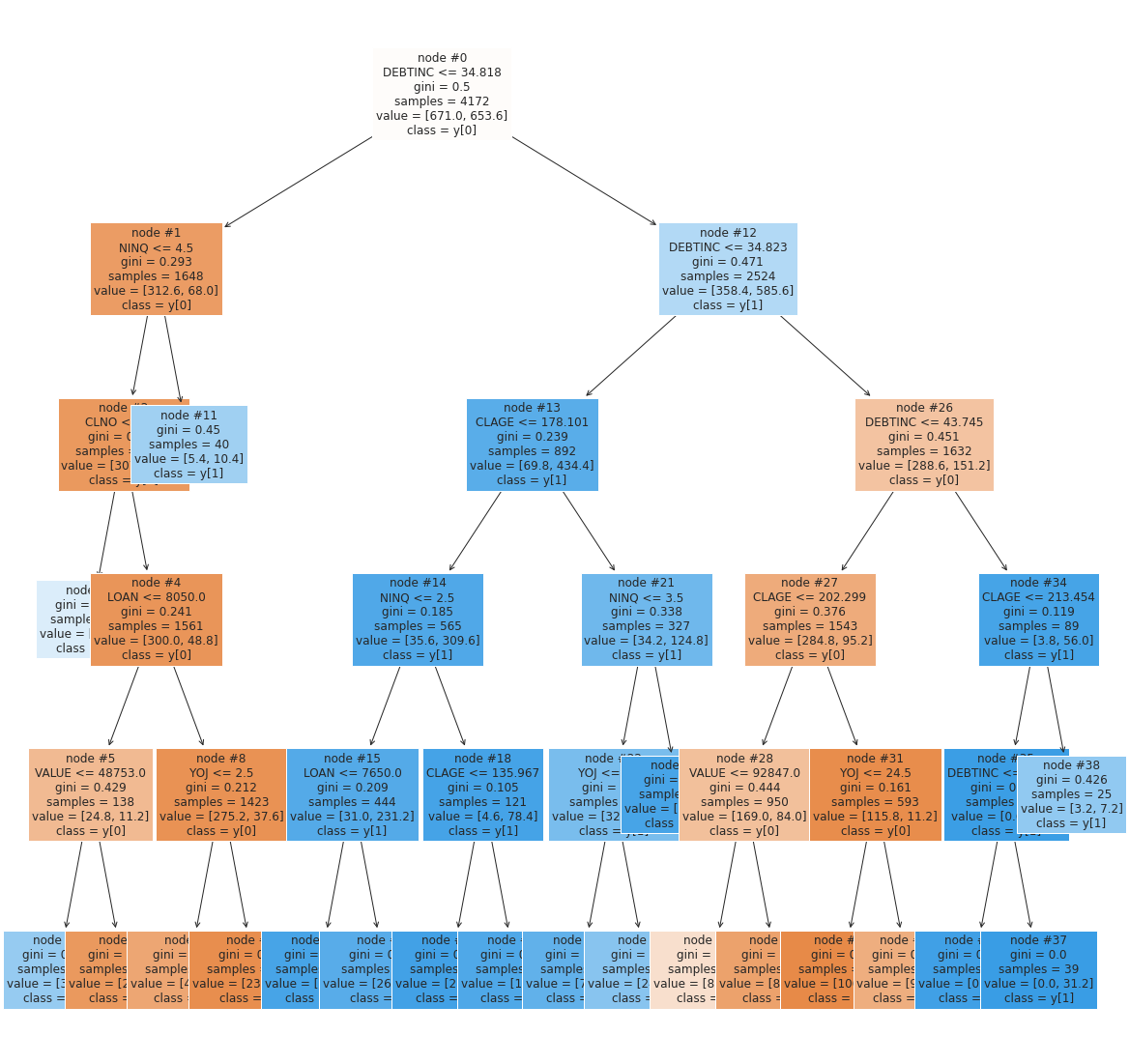
Observations
- The first split is made on
DEBTINC(Debt-to-income ratio) which means it has a high ability to predict defaulters - Higher a
DEBTINCindicates the applicant is more likely to default (As seen with our observations in EDA)
The next high priority splits are made on:
NINQ(Number of recent credit inquiries) - Where a higherNINQis more likely to defaultCLAGE(Age of the oldest credit line in months) - Where a lowerCLAGEis more likely to default- These where not apparent as strong indicators in EDA indicating that many indicators fall in the scope of
DEBTINC, whereNINQandCLAGEhave less overlap.
Plotting Feature Importance
# Plotting the feature importance
importances = d_tree_tuned.feature_importances_
indices = np.argsort(importances)
plt.figure(figsize = (10, 10))
plt.title('Feature Importances')
plt.barh(range(len(indices)), importances[indices], color = 'orange', align = 'center')
plt.yticks(range(len(indices)), [features[i] for i in indices])
plt.xlabel('Relative Importance')
plt.show()
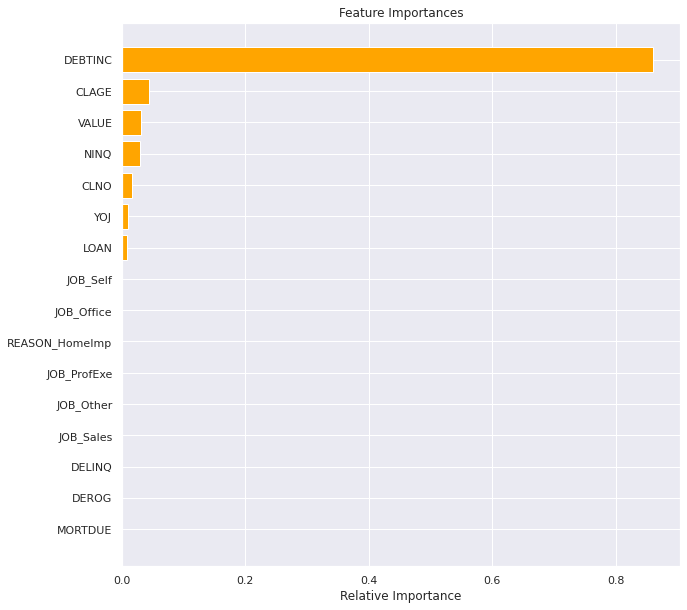
Observations
- Highlights just how much of a dominant feature
DEBTINCis compared to the others. CLAGE(age of oldest credit line in months),VALUE(Current value of the property) andNINQ(number of recent credit inquiries) are also a impactfull feature- Where lower
VALUEis more likely to default - Where lower
CLAGEis more likely to default - Where higher
NINQis more likely to default -
CLNO(number of existing credit lines) ,YOJ(years at present job)LOAN(loan amount), are the only other features that have an impact. - Surprising that
DEROG(number of serious delinquency or late payments) andDELINQ(number of delinquent credit lines) did not show any impact given our EDA suggesting they might. - Surprising that JOB_SALES and JOBS_SELF did not show any impact given our EDA suggesting they might.
Building a Random Forest Classifier
Random Forest is a bagging algorithm where the base models are Decision Trees. Samples are taken from the training data and on each sample a decision tree makes a prediction.
The results from all the decision trees are combined together and the final prediction is made using voting or averaging.
# Defining Random forest CLassifier
rf_base = RandomForestClassifier(random_state=7,criterion="entropy")
rf_base.fit(X_train,y_train)
RandomForestClassifier(criterion='entropy', random_state=7)
Checking the performance on the train dataset
#Checking performance on the training data
y_pred_train3 = rf_base.predict(X_train)
metrics_score(y_train,y_pred_train3)
precision recall f1-score support
0 1.00 1.00 1.00 3355
1 1.00 1.00 1.00 817
accuracy 1.00 4172
macro avg 1.00 1.00 1.00 4172
weighted avg 1.00 1.00 1.00 4172

Checking the performance on the test dataset
# Checking performance on the test data
y_pred_test3 = rf_base.predict(X_test)
metrics_score(y_test, y_pred_test3)
precision recall f1-score support
0 0.90 0.97 0.93 1416
1 0.85 0.59 0.69 372
accuracy 0.89 1788
macro avg 0.87 0.78 0.81 1788
weighted avg 0.89 0.89 0.88 1788

Add data to results table
# Adding the results to the table
new_row = {'Model_Name': 'random_forest',
'Train_f1': 100,
'Train_recall': 100,
'Test_f1': 69,
'Test_recall': 59,
'Test_precision': 85}
results = pd.concat([results, pd.DataFrame([new_row])], ignore_index=True)
# Print the updated DataFrame
print(results)
Model_Name Train_f1 Train_recall Test_f1 Test_recall Test_precision
0 d_tree_base 100 100 60 56 64
1 d_tree_base 68 79 67 75 60
2 random_forest 100 100 69 59 85
Observations
- The base random forest model is highly overfitted as expected
- It is naturally favoring precision, the oposite of what we want
Random Forest with class weights
# Defining Random Forest model with class weights class_weight={0: 0.2, 1: 0.8}
rf_weighted = RandomForestClassifier(random_state = 7, class_weight = {0: 0.2, 1: 0.8})
# Fitting Random Forest model
rf_weighted.fit(X_train,y_train)
RandomForestClassifier(class_weight={0: 0.2, 1: 0.8}, random_state=7)
Checking the performance on the train dataset
# Checking performance on the train data
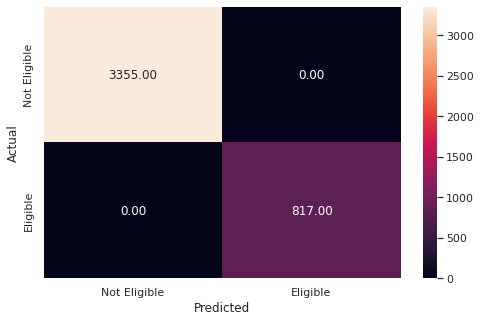
y_pred_train4 = rf_weighted.predict(X_train)
metrics_score(y_train,y_pred_train4)
precision recall f1-score support
0 1.00 1.00 1.00 3355
1 1.00 1.00 1.00 817
accuracy 1.00 4172
macro avg 1.00 1.00 1.00 4172
weighted avg 1.00 1.00 1.00 4172

Checking the performance on the test dataset
# Checking performance on the test data
y_pred_test4 = rf_weighted.predict(X_test)
metrics_score(y_test, y_pred_test4)
precision recall f1-score support
0 0.89 0.98 0.93 1416
1 0.87 0.55 0.68 372
accuracy 0.89 1788
macro avg 0.88 0.76 0.80 1788
weighted avg 0.89 0.89 0.88 1788

# Adding the results to the table
new_row = {'Model_Name': 'random_forest',
'Train_f1': 100,
'Train_recall': 100,
'Test_f1': 68,
'Test_recall': 55,
'Test_precision': 87}
results = pd.concat([results, pd.DataFrame([new_row])], ignore_index=True)
# Print the updated DataFrame
print(results)
Model_Name Train_f1 Train_recall Test_f1 Test_recall Test_precision
0 d_tree_base 100 100 60 56 64
1 d_tree_base 68 79 67 75 60
2 random_forest 100 100 69 59 85
3 random_forest 100 100 68 55 87
Weighting the random forest has dropped both the f1-score and recall scores
Random Forest Classifier Hyperparameter Tuning
# Choose the type of classifier
rf_tuned = RandomForestClassifier(criterion = "entropy", random_state = 7)
# Grid of parameters to choose from
parameters = {"n_estimators": [100, 110],
"max_depth": [5,6],
"max_leaf_nodes": [8,10],
"min_samples_split":[20],
'criterion': ['gini'],
"max_features": ['sqrt'],
"class_weight": ["balanced",{0: 0.2, 1: 0.8}]
}
# Type of scoring used to compare parameter combinations - recall score for class 1
scorer = metrics.make_scorer(recall_score, pos_label = 1)
# Run the grid search on the training data using scorer=scorer and cv=5
grid_obj = GridSearchCV(rf_tuned, parameters, scoring = scorer, cv = 5)
grid_obj = grid_obj.fit(X_train, y_train)
# Save the best estimator to variable rf_estimator_tuned
rf_tuned = grid_obj.best_estimator_
#Fit the best estimator to the training data
rf_tuned.fit(X_train, y_train)
RandomForestClassifier(class_weight='balanced', max_depth=6,
max_features='sqrt', max_leaf_nodes=10,
min_samples_split=20, random_state=7)
Checking the performance on the train dataset
# Checking performance on the training data
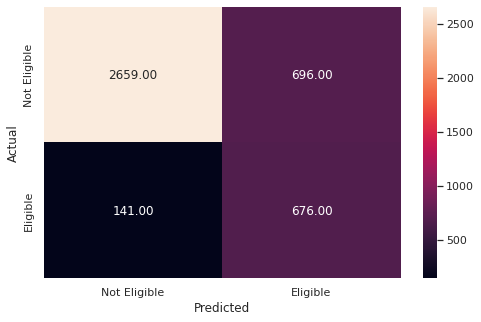
y_pred_train5 = rf_tuned.predict(X_train)
metrics_score(y_train, y_pred_train5)
precision recall f1-score support
0 0.95 0.79 0.86 3355
1 0.49 0.83 0.62 817
accuracy 0.80 4172
macro avg 0.72 0.81 0.74 4172
weighted avg 0.86 0.80 0.82 4172

Checking the performance on the test dataset
# Checking performace on test dataset
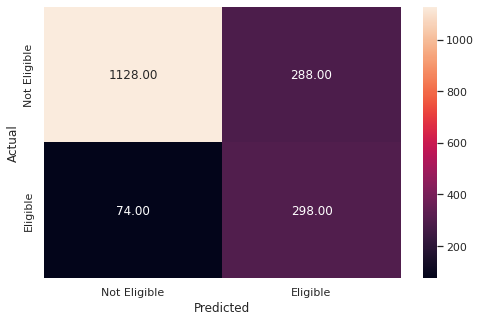
y_pred_test5 = rf_tuned.predict(X_test)
metrics_score(y_test, y_pred_test5)
precision recall f1-score support
0 0.94 0.80 0.86 1416
1 0.51 0.80 0.62 372
accuracy 0.80 1788
macro avg 0.72 0.80 0.74 1788
weighted avg 0.85 0.80 0.81 1788

# Adding the results to the table
new_row = {'Model_Name': 'random_forest',
'Train_f1': 62,
'Train_recall': 83,
'Test_f1': 62,
'Test_recall': 80,
'Test_precision': 51}
results = pd.concat([results, pd.DataFrame([new_row])], ignore_index=True)
# Print the updated DataFrame
print(results)
results.to_csv('results.csv')
Model_Name Train_f1 Train_recall Test_f1 Test_recall Test_precision
0 d_tree_base 100 100 60 56 64
1 d_tree_base 68 79 67 75 60
2 random_forest 100 100 69 59 85
3 random_forest 100 100 68 55 87
4 random_forest 62 83 62 80 51
Observations
- The model is no longer overfitted with a equal training and testing f1-score of 62%.
- After multiple tunnings, we can conclude that the limited dataset makes it difficult to achieve a high accuracy
- However was able to tune the model to maximize recall and achieve a score of 80%.
Plot the Feature importance of the tuned Random Forest
# importance of features in the random forest
importances = rf_tuned.feature_importances_
indices = np.argsort(importances)
feature_names = list(X.columns)
plt.figure(figsize = (12, 12))
plt.title('Feature Importances')
plt.barh(range(len(indices)), importances[indices], color = 'red', align = 'center')
plt.yticks(range(len(indices)), [feature_names[i] for i in indices])
plt.xlabel('Relative Importance')
plt.show()
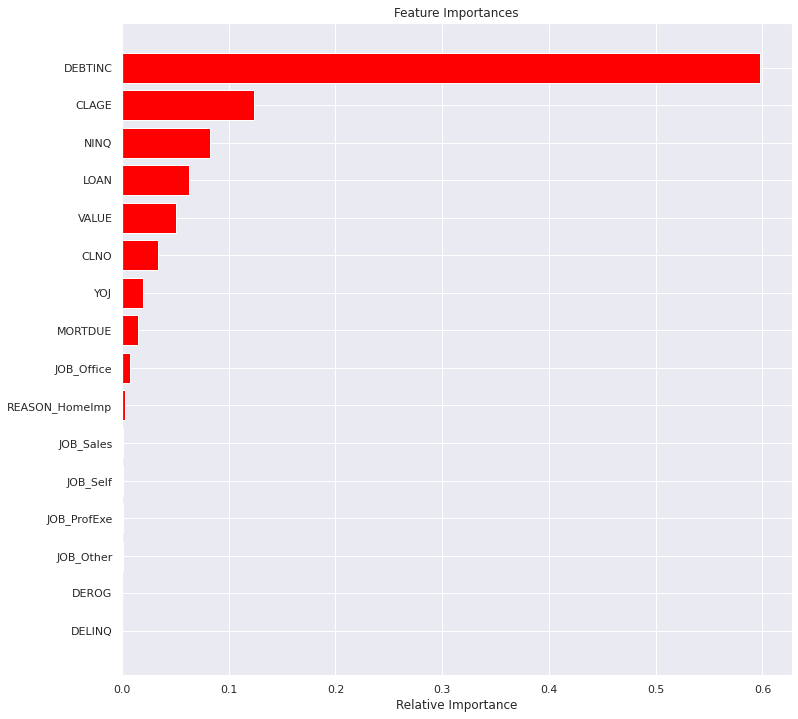
Observations
- Random forest feature importance confirms many of the same features before.
DEBTINCcontinues to be a dominant feature- CLAGE, VALUE and NINQ continue to show relevant importance
- Loan has increased in importance where a smaller loan is more likely to default
Conclusion and Recommendations
Goal
- Build a classification model to predict the customers who are likely to default on their loan
Process
-
Maximize Recall (false negatives) as banks lose more money when a customer defaults on thier loans.
-
Maintain a high F1-Score (overall accuracy) as false positives would result in a loss of interest profit for the bank.
-
Give recommendations to the bank on the important customer characteristics to consider while approving a loan.
Conclusions:
- As the models were tuned it was noticeable that the size (5960 data points) and composition (80/20 non-defaulters to defaulters) of the data was contributing to lower that ideal accuracy scores.
- Therefore, the focus was only on maximizing recall even at the expense of overall accuracy. (The F1-score was not as high as preferred).
We built and tuned two models to predict if a customer would likely default on their loan. The results of the two models are:
Tuned Decision Tree Model - F1-score 67% and recall 75% Hyper Tuned Random Forest Model - F1-score 62% and recall 80%
- The models are similarly balanced with 67% and 62% F1-scores.
- However, since the goal is to maximize recall the Hyper Tuned Random Forest Model is recommended.
- The recall was higher at 80% with an accuracy of 80%.
- This was achieved with a smaller data set and challenge of missing values.
Major Features (Characteristics)
The top two features identified by both the Decision Tree and the Random Forest Models
- Debt to income ratio (
DEBTINC) is a very powerful tool in predicting defaulters.- The Debt to income ratio was the top feature identified in both models.
- The higher a customers debt-to-income ratio, the more likely the customer is to default.
- Therefore the bank should use the debt-to-income ratio as the first indicator when evaluating a customer.
- The credit age (
CLAGE) of a customer is a feature to also review.- The credit age was the second most important feature identified in both models.
- The amount of time someone has had credit the better the bank can gauge how well they will repay their loans.
- Additional features identified by both models
VALUEandNINQwere both identified in the top 5 for both models, however, they were not ranked of the same importance.VALUEwas 3rd for the Decision Tree Model and 5th for the Random Forest Model.NINQwas 4th for the Decision Tree Model and 3rd for the Random Forest Model.
- Feature identified by only one of the models in the top 5
CLNOwas 5th for the Decision Tree ModelLOANwas 4th for the Random Forest Model
- Derogatory report (DEROG) and delinquency credit report (DELINQ) are also very important indicators
- Customers with DEROG >= 7 or DELINQ >= 6 all defauted on their loans.
- Both of these were identified during the EDA.
- Self-employed customers have a higher probability of defaulting on their loans.
- Identified during the EDA
Overall
-
More data should be used to create a stronger model.
- More care should be taken when gathering and inputting data.
-
There were possible errors and a large anount of missing data that could have assisted with improving the overall model.
DEBTINChad the most missing values and had some possible inaccuracies in the data. This feature was found to be the MOST important feature for predicting if a person would default. The importance could be because of the correction of the missing values or it could be because it is important. This needs to be evaluated further.